The effects of missing data
Robert W Schlegel
2019-11-06
Last updated: 2019-11-06
workflowr checks: (Click a bullet for more information)-
✔ R Markdown file: up-to-date
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
-
✔ Environment: empty
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
-
✔ Seed:
set.seed(666)The command
set.seed(666)was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible. -
✔ Session information: recorded
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
-
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.✔ Repository version: ff0bb95
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can usewflow_publishorwflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.Ignored files: Ignored: .Rhistory Ignored: .Rproj.user/ Ignored: data/global/ Ignored: data/global_results.Rda Ignored: data/global_test_trend.Rda Ignored: data/global_var_trend.Rda Ignored: data/global_var_trend_old.Rda Ignored: data/random_bp_results_100.Rda Ignored: data/random_bp_results_1000.Rda Ignored: data/random_results_100.Rda Ignored: data/random_results_1000.Rda Ignored: data/sst_ALL_bp_results.Rda Untracked files: Untracked: analysis/WA_pixels.Rda Untracked: analysis/WA_pixels_res.Rda Untracked: docs/figure/time_series_length.Rmd/ Unstaged changes: Modified: .DS_Store Modified: .Rprofile Modified: .gitignore Modified: CODE_OF_CONDUCT.md Modified: LICENSE Modified: LICENSE.md Modified: LaTeX/FMars.csl Modified: LaTeX/Frontiers_Template.docx Modified: LaTeX/MHWdetection.docx Modified: LaTeX/MHWdetection.tex Modified: LaTeX/PDF examples/frontiers.pdf Modified: LaTeX/PDF examples/frontiers_SupplementaryMaterial.pdf Modified: LaTeX/README Modified: LaTeX/Supplementary_Material.docx Modified: LaTeX/YM-logo.eps Modified: LaTeX/fig_1.jpg Modified: LaTeX/fig_1.pdf Modified: LaTeX/fig_1_flat.jpg Modified: LaTeX/fig_1_flat.pdf Modified: LaTeX/fig_2.jpg Modified: LaTeX/fig_2.pdf Modified: LaTeX/fig_3.jpg Modified: LaTeX/fig_3.pdf Modified: LaTeX/fig_4.jpg Modified: LaTeX/fig_4.pdf Modified: LaTeX/fig_5.jpg Modified: LaTeX/fig_5.pdf Modified: LaTeX/fig_6.jpg Modified: LaTeX/fig_6.pdf Modified: LaTeX/fig_S1.jpg Modified: LaTeX/fig_S1.pdf Modified: LaTeX/fig_S2.jpg Modified: LaTeX/fig_S2.pdf Modified: LaTeX/fig_S3.jpg Modified: LaTeX/fig_S3.pdf Modified: LaTeX/fig_S4.jpg Modified: LaTeX/fig_S4.pdf Modified: LaTeX/fig_S5.jpg Modified: LaTeX/fig_S5.pdf Modified: LaTeX/figures.zip Modified: LaTeX/frontiers.tex Modified: LaTeX/frontiersFPHY.cls Modified: LaTeX/frontiersHLTH.cls Modified: LaTeX/frontiersSCNS.cls Modified: LaTeX/frontiersSCNS.log Modified: LaTeX/frontiers_SupplementaryMaterial.tex Modified: LaTeX/frontiers_suppmat.cls Modified: LaTeX/frontiersinHLTH&FPHY.bst Modified: LaTeX/frontiersinSCNS_ENG_HUMS.bst Modified: LaTeX/logo1.eps Modified: LaTeX/logo1.jpg Modified: LaTeX/logo2.eps Modified: LaTeX/logos.eps Modified: LaTeX/logos.jpg Modified: LaTeX/stfloats.sty Modified: LaTeX/table_1.xlsx Modified: LaTeX/table_2.xlsx Modified: LaTeX/test.bib Modified: MHWdetection.Rproj Modified: TODO Modified: _references/1-s2.0-S0921818106002736-main.pdf Modified: _references/1-s2.0-S092181810600275X-main.pdf Modified: _references/1-s2.0-S0921818106002761-main.pdf Modified: _references/1-s2.0-S0921818106002852-main.pdf Modified: _references/1405.3904.pdf Modified: _references/1520-0450%282001%29040%3C0762%3Aotdoah%3E2.0.co%3B2.pdf Modified: _references/2013_Extremes_Workshop_Report.pdf Modified: _references/24868781.pdf Modified: _references/24870362.pdf Modified: _references/26192647.pdf Modified: _references/994.full.pdf Modified: _references/A_1019841717369.pdf Modified: _references/Banzon et al 2014.pdf Modified: _references/Brown_et_al-2008-Journal_of_Geophysical_Research%3A_Atmospheres_%281984-2012%29.pdf Modified: _references/Different_ways_to_compute_temperature_re.pdf Modified: _references/Gilleland et al 2013.pdf Modified: _references/Gilleland_2006.pdf Modified: _references/Kuglitsch_et_al-2010-Geophysical_Research_Letters.pdf Modified: _references/Modeling Waves of Extreme Temperature The Changing Tails of Four Cities.pdf Modified: _references/Normals-Guide-to-Climate-190116_en.pdf Modified: _references/Reynolds et al 2007.pdf Modified: _references/Risk_of_Extreme_Events_Under_Nonstationa.pdf Modified: _references/Russo_et_al-2014-Journal_of_Geophysical_Research%3A_Atmospheres.pdf Modified: _references/WCDMP_72_TD_1500_en__1.pdf Modified: _references/WMO 49 v1 2015.pdf Modified: _references/WMO No 1203.pdf Modified: _references/WMO-TD No 1377.pdf Modified: _references/WMO_100_en.pdf Modified: _references/bams-d-12-00066.1.pdf Modified: _references/c058p193.pdf Modified: _references/cc100.pdf Modified: _references/clivar14.pdf Modified: _references/coles1994.pdf Modified: _references/ecology.pdf Modified: _references/joc.1141.pdf Modified: _references/joc.1432.pdf Modified: _references/returnPeriod.pdf Modified: _references/s00382-014-2287-1.pdf Modified: _references/s00382-014-2345-8.pdf Modified: _references/s00382-015-2638-6.pdf Modified: _references/s10584-006-9116-4.pdf Modified: _references/s10584-007-9392-7.pdf Modified: _references/s10584-010-9944-0.pdf Modified: _references/s10584-012-0659-2.pdf Modified: _references/s10584-014-1254-5.pdf Modified: _references/s13253-013-0161-y.pdf Modified: _references/wcrpextr.pdf Modified: _workflowr.yml Modified: analysis/Climatologies_and_baselines.Rmd Modified: analysis/Short_climatologies.Rmd Modified: analysis/about.Rmd Modified: analysis/bibliography.bib Modified: analysis/gridded_products.Rmd Modified: analysis/r_vs_python_arguments.Rmd Modified: analysis/variance.Rmd Modified: code/README.md Modified: data/.gitignore Modified: data/best_table_average.Rda Modified: data/best_table_focus.Rda Modified: data/python/clim_py.csv Modified: data/python/clim_py_joinAG_1.csv Modified: data/python/clim_py_joinAG_5.csv Modified: data/python/clim_py_joinAG_no.csv Modified: data/python/clim_py_minD_3.csv Modified: data/python/clim_py_minD_7.csv Modified: data/python/clim_py_pctile_80.csv Modified: data/python/clim_py_pctile_95.csv Modified: data/python/clim_py_pctile_99.csv Modified: data/python/clim_py_random.csv Modified: data/python/clim_py_spw_11.csv Modified: data/python/clim_py_spw_51.csv Modified: data/python/clim_py_spw_no.csv Modified: data/python/clim_py_whw_3.csv Modified: data/python/clim_py_whw_7.csv Modified: data/python/mhwBlock.csv Modified: data/python/mhws_py.csv Modified: data/python/mhws_py_joinAG_1.csv Modified: data/python/mhws_py_joinAG_5.csv Modified: data/python/mhws_py_joinAG_no.csv Modified: data/python/mhws_py_minD_3.csv Modified: data/python/mhws_py_minD_7.csv Modified: data/python/mhws_py_pctile_80.csv Modified: data/python/mhws_py_pctile_95.csv Modified: data/python/mhws_py_pctile_99.csv Modified: data/python/mhws_py_random.csv Modified: data/python/mhws_py_spw_11.csv Modified: data/python/mhws_py_spw_51.csv Modified: data/python/mhws_py_spw_no.csv Modified: data/python/mhws_py_whw_3.csv Modified: data/python/mhws_py_whw_7.csv Modified: data/python/sst_WA.csv Modified: data/python/sst_WA_miss_ice.csv Modified: data/python/sst_WA_miss_random.csv Modified: data/sst_ALL_results.Rda Modified: data/table_1.csv Modified: data/table_2.csv Modified: docs/portrait.pdf Modified: output/README.md Modified: output/effect_event.pdf Modified: output/fig_2_missing_only.pdf Modified: output/stitch_plot_WA.pdf Modified: output/stitch_sub_plot_WA.pdf Modified: poster/Figures/CSIRO_logo.jpeg Modified: poster/Figures/Dal_logo.jpg Modified: poster/Figures/all_logo_long.jpg Modified: poster/Figures/all_logos.jpg Modified: poster/Figures/logo_stitching.odp Modified: poster/Figures/ofi_logo.jpg Modified: poster/Figures/uwc-logo.jpg Modified: poster/MHWdetection.bib Modified: poster/MyBib.bib Modified: poster/landscape.Rmd Modified: poster/landscape.pdf Modified: poster/portrait.Rmd Modified: poster/portrait.pdf
Expand here to see past versions:
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | ff0bb95 | robwschlegel | 2019-11-06 | Publish the sub-optimal test vignettes |
| Rmd | 61bf228 | robwschlegel | 2019-11-06 | Prepped trend vignette for site re-build |
| Rmd | efa959e | robwschlegel | 2019-11-06 | Prepped missing data vignette for website re-build |
| Rmd | 158aa0b | robwschlegel | 2019-05-06 | Updated project interface |
| html | 158aa0b | robwschlegel | 2019-05-06 | Updated project interface |
| html | 38559da | robwschlegel | 2019-03-19 | Build site. |
| Rmd | 970b22c | robwschlegel | 2019-03-19 | Publish the vignettes from when this was a pkgdown framework |
| html | fa7fd57 | robwschlegel | 2019-03-19 | Build site. |
| Rmd | 64ac134 | robwschlegel | 2019-03-19 | Publish analysis files |
Overview
The purpose of this vignette is to briefly show the process used to go about quantify the effects that missing data have on the detection of MHWs. Specifically, the relationship between the percentage of missing data and how the MHW metrics may differ from those detected against the same time series with no missing data.
The missing data will be ‘created’ by striking out existing data from the three pre-packaged time series in the heatwaveR package, which themselves have no missing data. These data will first be detrended so that the random missing data removed will not be conflated with any trend in the data.
# The packages used in this analysis
library(tidyverse)
library(heatwaveR)
library(lubridate)
library(ncdf4)
library(doParallel)
# The custom functions written for the analysis
source("code/functions.R")# The function used to de-trend the time series:
detrend <- function(df){
resids <- broom::augment(lm(temp ~ t, df))
res <- df %>%
mutate(temp = round((temp - resids$.fitted),2))
return(res)
}The effects that missing data have on the results are investigated with the same methodology as the time series length and long-term trend tests, with the difference that data are being randomly removed before the analysis is run.
Random missing data
First up we begin with the random removal of increasing proportions of the data. We are going to use the full 37 year time series for these missing data experiments. We will randomly remove 0 – 50% of the data from each of the three times series in 1% steps. This is not being repeated here so the results will look very jagged. The necessary replication of this study is performed by repeating it on 1000 randomly selected pixels, which may be seen in ‘code/workflow.R’ in the GitHub repository.
control_missing <- function(prop, df){
# NB: Don't allow sampling of first and last value to ensure
# all time series are the same length
ts_length <- nrow(df)
miss_index <- sample(seq(2, ts_length-1, 1), ts_length*prop, replace = F)
res <- df %>%
mutate(row_index = 1:n(),
temp = replace(temp, which(row_index %in% miss_index), NA),
test = "missing",
index_vals = prop) %>%
dplyr::select(test, index_vals, t, temp)
return(res)
}Analysis
The full analysis on the results, including the functions shown above, is run for all of the tests (time series length, missing data, and long-term trend) all at once to ensure consistency of methodology across tests. For this reason the exact step-by-step process for the missing data analysis is not laid out below. To investigate the source code one may go to ‘code/workflow.R’ in the GitHub repository. A link to that site may be found in the top right of the navigation bar for this site (the GitHub icon).
sst_ALL <- rbind(mutate(sst_WA, site = "WA"),
mutate(sst_NW_Atl, site = "NW_Atl"),
mutate(sst_Med, site = "Med"))
system.time(
sst_ALL_results <- plyr::ddply(sst_ALL, c("site"), single_analysis, .parallel = T,
full_seq = T, clim_metric = F, count_miss = T, windows = T)
) # 65 secondsResults
A more thorough explanation of the results may be found in the manuscript. Below we show what the simple results calculated above for the effect of missing data on the count of the focal MHW look like.
sst_ALL_results %>%
filter(test == "missing",
var == "focus_count",
id == "mean_perc") %>%
ggplot(aes(x = index_vals*100, y = val, colour = site)) +
geom_line() +
labs(x = "Missing data (%)", y = "Change from control (% count of MHWs)", colour = "Site")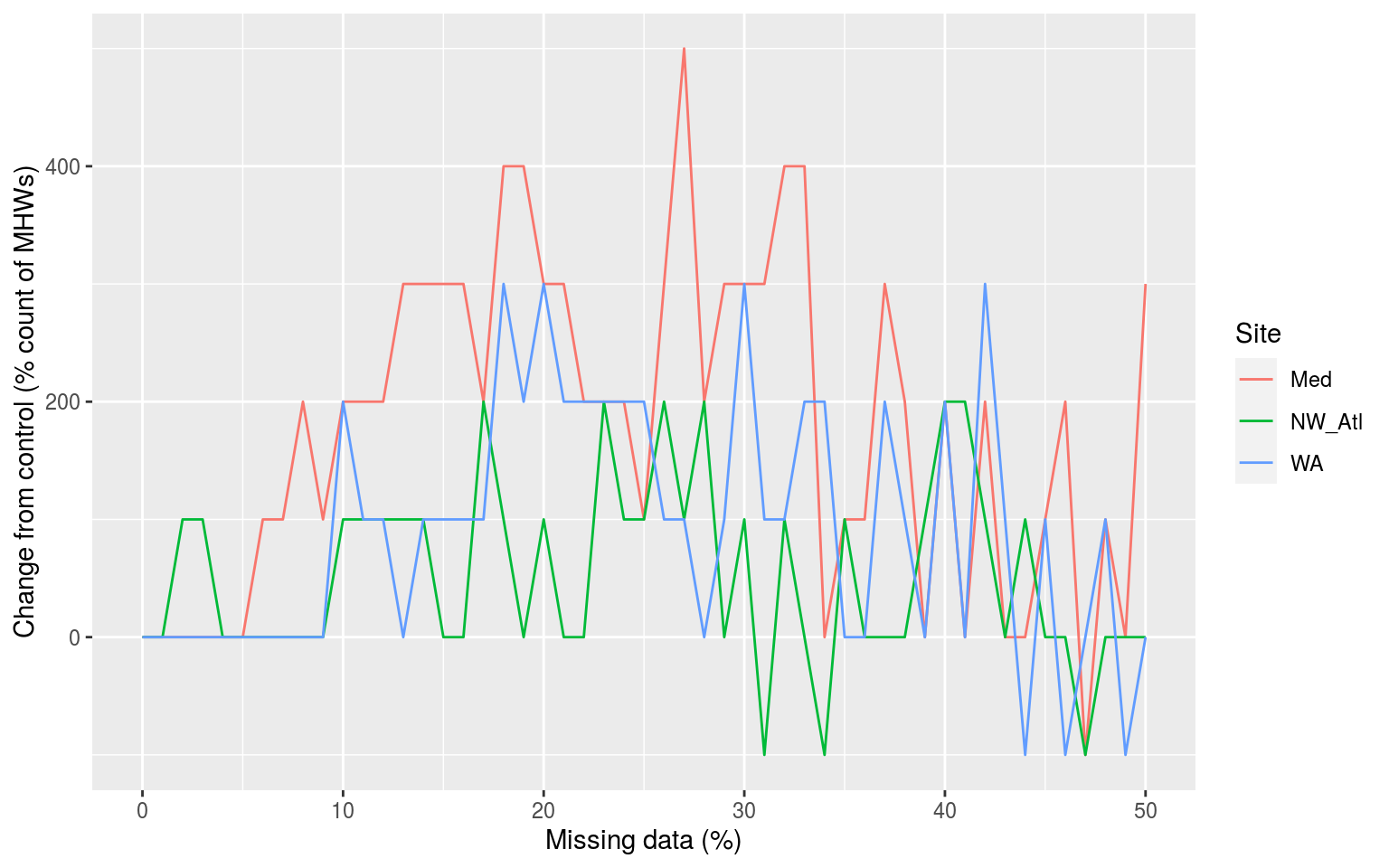
References
Session information
sessionInfo()R version 3.6.1 (2019-07-05)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Ubuntu 16.04.6 LTS
Matrix products: default
BLAS: /usr/lib/openblas-base/libblas.so.3
LAPACK: /usr/lib/libopenblasp-r0.2.18.so
locale:
[1] LC_CTYPE=en_CA.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_CA.UTF-8 LC_COLLATE=en_CA.UTF-8
[5] LC_MONETARY=en_CA.UTF-8 LC_MESSAGES=en_CA.UTF-8
[7] LC_PAPER=en_CA.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_CA.UTF-8 LC_IDENTIFICATION=C
attached base packages:
[1] parallel stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] doParallel_1.0.15 iterators_1.0.10 foreach_1.4.4
[4] ncdf4_1.17 lubridate_1.7.4 heatwaveR_0.4.1.9003
[7] forcats_0.4.0 stringr_1.4.0 dplyr_0.8.3
[10] purrr_0.3.3 readr_1.3.1 tidyr_1.0.0
[13] tibble_2.1.3 ggplot2_3.2.1.9000 tidyverse_1.2.1
loaded via a namespace (and not attached):
[1] tidyselect_0.2.5 xfun_0.10 haven_2.1.1
[4] lattice_0.20-35 colorspace_1.4-1 vctrs_0.2.0
[7] generics_0.0.2 viridisLite_0.3.0 htmltools_0.4.0
[10] yaml_2.2.0 plotly_4.9.0 rlang_0.4.1
[13] R.oo_1.22.0 pillar_1.4.2 glue_1.3.1
[16] withr_2.1.2 R.utils_2.7.0 modelr_0.1.5
[19] readxl_1.3.1 lifecycle_0.1.0 munsell_0.5.0
[22] gtable_0.3.0 workflowr_1.1.1 cellranger_1.1.0
[25] rvest_0.3.4 R.methodsS3_1.7.1 htmlwidgets_1.5.1
[28] codetools_0.2-15 evaluate_0.14 labeling_0.3
[31] knitr_1.25 broom_0.5.2 Rcpp_1.0.2
[34] backports_1.1.5 scales_1.0.0 jsonlite_1.6
[37] hms_0.5.1 digest_0.6.22 stringi_1.4.3
[40] grid_3.6.1 rprojroot_1.3-2 cli_1.1.0
[43] tools_3.6.1 maps_3.3.0 magrittr_1.5
[46] lazyeval_0.2.2 crayon_1.3.4 whisker_0.4
[49] pkgconfig_2.0.3 zeallot_0.1.0 data.table_1.12.6
[52] xml2_1.2.2 assertthat_0.2.1 rmarkdown_1.16
[55] httr_1.4.1 rstudioapi_0.10 R6_2.4.0
[58] nlme_3.1-137 git2r_0.23.0 compiler_3.6.1 This reproducible R Markdown analysis was created with workflowr 1.1.1